

Description
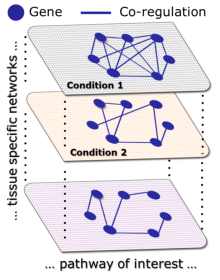
Our working hypothesis is that genes belonging to a condition-specific pathway are actively co-regulated only in specific conditions when the pathway is active, but not in others, independently of their absolute level of expression. We developed a network-based algorithm, DINA, which is able to identify sets of genes which are significantly co-regulated only in specific conditions, but not in others. The algorithm starts with a set of M genes ( i.e. genes belonging to the same pathway ) and a set of N networks ( i.e. the thirty tissue-specific gene networks ). It then computes a ''co-regulation probability'' for the M genes in each of the N networks; this probability is proportional to the number of edges among the genes in each of the networks. DINA then quantifies how variable the co-regulation probability is across the N networks. Variability is quantified using an entropy-based measure (H).
How to cite DINA: Gambardella, G. et al., Differential network analysis for the identification of condition-specific pathway activity and regulation. Bioinformatics, 2013 - Abstract
probability is across the N networks. Variability is quantified using an entropy-based measure (H).
How to cite DINA: Gambardella, G. et al., Differential network analysis for the identification of condition-specific pathway activity and regulation. Bioinformatics, 2013 - Abstract



© Telethon Institute of Genetics and Medicine







Insert the list of genes (i.e. Official Gene Symbols)



